
2018–02–20 Update: Adds two images (random forest and gradient boosting).
2019–05–25 Update: I’ve published a post covering another importance measure — SHAP values — on my personal blog and on Medium.
This post is inspired by a Kaggle kernel and its discussions [1]. I’d like to do a brief review of common algorithms to measure feature importance with tree-based models. We can interpret the results to check intuition(no surprisingly important features), do feature selection, and guide the direction of feature engineering.
Here’s the list of measures we’re going to cover with their associated models:
-
Random Forest: Gini Importance or Mean Decrease in Impurity (MDI) [2]
-
Random Forest: Permutation Importance or Mean Decrease in Accuracy (MDA) [2]
-
Random Forest: Boruta [3]
-
Gradient Boosting: Split-based and Gain-based Measures.
Note that measure 2 and 3 are theoretically applicable to all tree-based models.
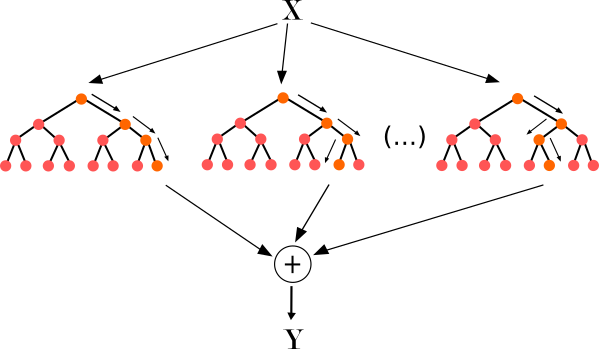
Random Forest (Source)
Gini Importance / Mean Decrease in Impurity (MDI)
According to [1], MDI counts the times a feature is used to split a node, weighted by the number of samples it splits:
Gini Importance or Mean Decrease in Impurity (MDI) calculates each feature importance as the sum over the number of splits (across all tress) that include the feature, proportionally to the number of samples it splits.
However, Gilles Louppe gave a different version in [4]. Instead of counting splits, the actual decrease in node impurity is summed and averaged across all trees. (weighted by the number of samples it splits).
In scikit-learn, we implement the importance as described in [1] (often cited, but unfortunately rarely read…). It is sometimes called “gini importance” or “mean decrease impurity” and is defined as the total decrease in node impurity (weighted by the probability of reaching that node (which is approximated by the proportion of samples reaching that node)) averaged over all trees of the ensemble.
In R package randomForest, the implementation seems to be consistent with what Gilles Louppe described (another popular package, ranger, also seems to be doing the same) [5][6]:
The last column is the mean decrease in Gini index.
Finally, quoting the Element of Statistical Learning [7]:
At each split in each tree, the improvement in the split-criterion is the importance measure attributed to the splitting variable, and is accumulated over all the trees in the forest separately for each variable.
Permutation Importance or Mean Decrease in Accuracy (MDA)
This is IMO most interesting measure, because it is based on experiments on out-of-bag(OOB) samples, via destroying the predictive power of a feature without changing its marginal distribution. Because scikit-learn doesn’t implement this measure, people who only use Python may not even know it exists.
Here’s permutation importance described in the Element of Statistical Learning:
Random forests also use the OOB samples to construct a different variable-importance measure, apparently to measure the prediction strength of each variable. When the bth tree is grown, the OOB samples are passed down the tree, and the prediction accuracy is recorded. Then the values for the jth variable are randomly permuted in the OOB samples, and the accuracy is again computed. The decrease in accuracy as a result of this permuting is averaged over all trees, and is used as a measure of the importance of variable j in the random forest. … The randomization effectively voids the effect of a variable, much like setting a coefficient to zero in a linear model (Exercise 15.7). This does not measure the effect on prediction were this variable not available, because if the model was refitted without the variable, other variables could be used as surrogates.
For other tree models without bagging mechanism (hence no OOB), we can create a separate validation set and use it to evaluate the decrease in accuracy, or just use the training set [10].
This algorithm gave me an impression that it should be model-agnostic (can be applied on any classifier/regressors), but I’ve not seen literatures discussing its theoretical and empirical implications on other models. The idea to use it on neural networks was briefly mentioned on the Internet. And the same source claimed the algorithm works well on SVM models [8].
(20200908 Update: It’s been shown that permutation importance can be very misleading when the features are significantly correlated with each other. Please see my latest post for more information and a few alternative measures)
Boruta
Boruta is the name of an R package that implements a novel feature selection algorithm. It randomly permutes variables like Permutation Importance does, but performs on all variables at the same time and concatenates the shuffled features with the original ones. The concatenated result is used to fit the model.
Daniel Homola, who also wrote the Python version of Boruta(BorutaPy), gave an wonderful overview of the Boruta algorithm in his blog post [9].
The shuffled features (a.k.a. shadow features) are basically noises with identical marginal distribution w.r.t the original feature. We count the times a variable performs better than the “best” noise and calculate the confidence towards it being better than noise (the p-value) or not. Features which are confidently better are marked “confirmed”, and those which are confidently on par with noises are marked “rejected”. Then we remove those marked features and repeat the process until all features are marked or a certain number of iteration is reached.
Although Boruta is a feature selection algorithm, we can use the order of confirmation/rejection as a way to rank the importance of features.
Feature Importance Measure in Gradient Boosting Models
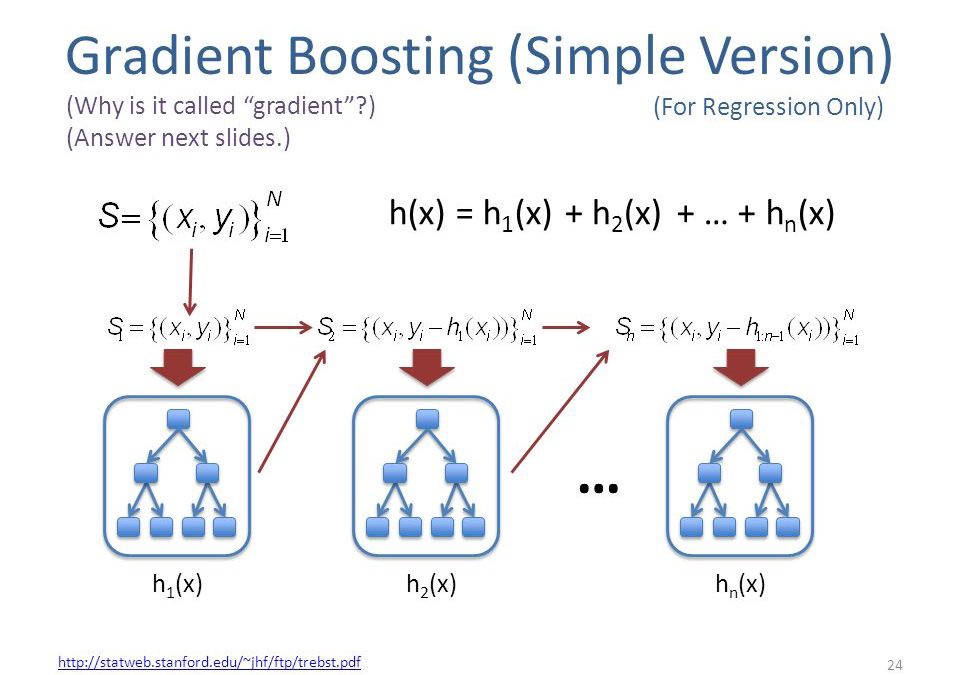
Gradient Boosting (Source)
For Kagglers, this part should be familiar due to the extreme popularity of XGBoost and LightGBM. Both packages implement more of the same measures (XGBoost has one more):
(LightGBM) importance_type (string, optional (default=”split”)) — How the importance is calculated. If “split”, result contains numbers of times the feature is used in a model. If “gain”, result contains total gains of splits which use the feature.
(XGBoost) ‘weight’ — the number of times a feature is used to split the data across all trees. ‘gain’ — the average gain of the feature when it is used in trees ‘cover’ — the average coverage of the feature when it is used in trees, where coverage is defined as the number of samples affected by the split
First measure is split-based and is very similar with the one given by [1] for Gini Importance. But it doesn’t take the number of samples into account.
The second measure is gain-based. It’s basically the same as the Gini Importance implemented in R packages and in scikit-learn with Gini impurity replaced by the objective used by the gradient boosting model.
The final measure, implemented exclusively in XGBoost, is counting the number of samples affected by the splits based on a feature.
The default measure of both XGBoost and LightGBM is the split-based one. I think this measure will be problematic if there are one or two feature with strong signals and a few features with weak signals. The model will exploit the strong features in the first few trees and use the rest of the features to improve on the residuals. The strong features will look not as important as they actually are. While setting lower learning rate and early stopping should alleviate the problem, also checking gain-based measure may be a good idea.
Note that these measures are purely calculated using training data, so there’s a chance that a split creates no improvement on the objective in the holdout set. This problem is more severe than in the random forest since gradient boosting models are more prone to over-fitting. It’s also one of the reason why I think Permutation Importance is worth exploring.
To Be Continued…
As usual, I will demonstrate some results of these measures on actual datasets in the next part.
Update: Part II has been published!
References
- Noise analysis of Porto Seguro’s features by olivier and its comment section. Kaggle.
- Alexis Perrie. Feature Importance in Random Forests.
- Miron B. Kursa, Witold R. Rudnicki (2010). Feature Selection with the Boruta Package. Journal of Statistical Software, 36(11) , p. 1–13.
- How are feature_importances in RandomForestClassifier determined? Stackoverflow.
- Andy Liaw. Pacakge ‘randomForest’. p. 17.
- Marvin N. Wright. Pacakge ‘ranger’. p. 14.
- Trevor Hastie, Robert Tibshirani, and Jerome Friedman. The Element of Statistical Learning. p. 593.
- Estimating importance of variables in a multilayer perceptron. CrossValidated.
- Daniel Homola. BorutaPy — an all relevant feature selection method.
- Hooker, G., & Mentch, L. (2019). Please Stop Permuting Features: An Explanation and Alternatives.